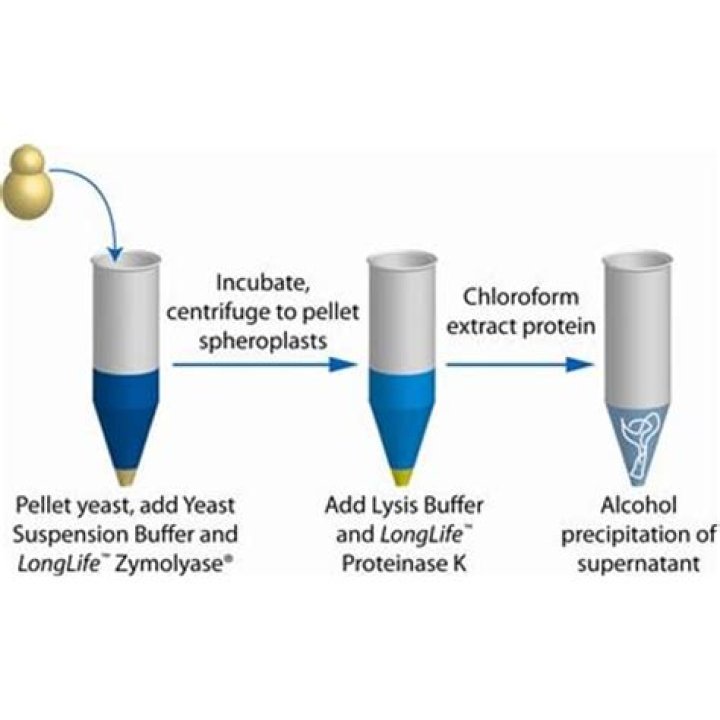
How do you isolate genomic DNA from yeast?
PROCEDURE
- Pick one yeast colony from the plate or spin down 100-200 μl of liquid yeast culture (OD600=0.4).
- Incubate for 5 minutes at 70°C.
- Add 300μl of 96-100 % ethanol, vortex.
- Spin down DNA and cell debris at 15 000 g for 3 minutes.
- Wash pellet with 70 % ethanol.
How do you PCR yeast?
Touch a yeast colony with a sterile 20µl pipet tip and put the cells directly into a PCR tube with PCR mix. The mix should be slightly cloudy but not dense with cells. Heat 8 minutes, 94 degrees (program 97 in the Robocycler). While they are heating, dilute the Taq polymerase.
Which method is used for separation of genomic DNA?
The DNA is then precipitated by adding isopropanol to the high-concentration salt solution. This forces the large genomic DNA molecules out of solution, while the smaller RNA fragments remain soluble. The insoluble DNA is then pelleted and separated from salt, isopropanol and RNA fragments via centrifugation.
Is yeast DNA circular or linear?
Yeast is a priviledged source of cytoplasmically replicating linear plasmids. These linear DNA molecules can be distinguished in a number of ways from the circular plasmids that replicate in the nucleus. One simple criterion is the large difference in sensitivity to ultraviolet light (UV) in vivo.
What is the meaning of genomic DNA?
Genomic deoxyribonucleic acid is chromosomal DNA, in contrast to extra-chromosomal DNAs like plasmids. It is also then abbreviated as gDNA. The genome of an organism (encoded by the genomic DNA) is the (biological) information of heredity which is passed from one generation of organism to the next.
What are the different methods of DNA extraction?
Some of the most common DNA extraction methods include organic extraction, Chelex extraction, and solid phase extraction. These methods consistently yield isolated DNA, but they differ in both the quality and the quantity of DNA yielded.
What is yeast colony?
When growing on solid surfaces, yeast, like other microorganisms, develops organized multicellular populations (colonies and biofilms) that are composed of differentiated cells with specialized functions.
What is the role of NaOH in DNA extraction?
NaOH helps to break down the cell wall, but more importantly, it disrupts the hydrogen bonding between the DNA bases, converting the double-stranded DNA (dsDNA) in the cell, including the genomic DNA (gDNA) and your plasmid, to single-stranded DNA (ssDNA).
What is a 260 280 ratio?
260/280 Ratio The ratio of absorbance at 260 nm and 280 nm is used to assess the purity of DNA and RNA. A ratio of ~1.8 is generally accepted as “pure” for DNA; a ratio of ~2.0 is generally accepted as “pure” for RNA.
What type of DNA is in yeast?
Electron microscopic analysis indicates that yeast nuclear DNA can be isolated as linear molecules ranging in size from 50 μm (1.2 × 108 daltons) to 355 μm (8.4 × 108 daltons). Analysis indicates the data is consistent with the hypothesis that each yeast chromosome contains a single, linear DNA duplex.
What is genomic DNA used for?
In research, genomic DNA are useful tools in applications such as PCR, library construction, Southern blotting, hybridizations, SNP analysis, and molecular diagnostic assays.
Is DNA extraction yeast PCR suitable for DNA sequencing?
DNA extracted by this method is suitable for a variety of PCR-based applications (including colony PCR, real-time qPCR, and DNA sequencing) for amplification of DNA fragments of ≤3500 bp. Keywords: DNA extraction yeast PCR
How do you extract DNA from yeast cells?
The protocol involves lysis of yeast colonies or cells from liquid culture in a lithium acetate (LiOAc)-SDS solution and subsequent precipitation of DNA with ethanol. Approximately 100 nanograms of total genomic DNA can be extracted from 1 × 10(7) cells.
How can we target genes in yeast using DNA modules?
Below, we describe basic methods for gene targeting in yeast using these modules, including design and production of PCR products for the construction of deletions, the introduction of mutations or epitope tags at the genomic locus and the swapping of tags and markers.
How to integrate PCR- derived DNA fragments with 40 bp of homology?
A high-efficiency transformation protocol is used for integration of PCR- derived DNA fragments with only 40 bp of homology to the target gene. Integration at the desired genomic locus can be verified by PCR using genomic DNA isolated in the rapid genomic DNA isolation protocol or by PCR of whole colonies.